How DNA Replicates
Matthew Meselson Franklin W. Stahl
 Matthew Meselson
Matthew Meselson
Matthew Meselson had a passion for physics and chemistry throughout his early life, often conducting science experiments in his family's garage. At the age of 16, he enrolled at the University of Chicago, beginning an academic career that led to doctoral studies at the California Institute of Technology under Linus Pauling. In addition to his widely known work demonstrating semi-conservative replication of DNA with Frank Stahl, Meselson has made many key discoveries in the molecular biology. He is also known for his work in limiting the proliferation of chemical and biological weapons. Meselson is a member of the National Academy of Science and a recipient of the Lasker Award. He continues to serve as a member of the faculty at Harvard University, where he has taught and conducted research since 1960.
 Franklin Stahl
Franklin Stahl
Following a sheltered life in a Boston suburb (Needham), Frank stumbled his way through college (Harvard, 1951) before fleeing to a graduate school in biology (U Rochester) to avoid the military draft. While in the graduate school, Frank took a course taught by A. H. (Gus) Doermann, and, for the first time in his life, he had a goal. With Gus, he studied genetic recombination in phage. To meet a departmental requirement, Frank took a summer course at Woods Hole, where he met Matt Meselson and began the work described in this Key Experiment. In 1959, Frank joined the faculty at the University of Oregon, Eugene, in their new Institute of Molecular Biology. He has been there ever since. Frank is now an emeritus faculty member who still enjoys teaching as well as family life and the natural wonders of Oregon.
What's the Big Deal?
Some experiments have proven so influential that they have been christened with the names of the scientists who performed them. The "Meselson–Stahl experiment" is one of those. It has also been called "the most beautiful experiment in biology," a title that has seemed to stick over the years. Why was the Meselson and Stahl experiment so important? Their experiment provided the first critical test of the Watson–Crick models for the structure of DNA and its replication, which were not universally accepted at the time. The convincing results of the Meselson–Stahl experiment, however, dispelled all doubts. DNA was no longer just an imaginary model; it was a real molecule, and its replication could be followed in the form of visually compelling bands in an ultracentrifuge. Meselson and Stahl found that these DNA bands behaved in the ultracentrifuge exactly as Watson and Crick postulated they should. Why was the Meselson–Stahl experiment "beautiful"? Because it was conceptually simple and yet sufficiently powerful to differentiate between several competing hypotheses for how DNA might replicate. Taken together, the Watson–Crick model and the Meselson–Stahl experiment marked the transition to the modern era of molecular biology, a turning point as impactful as the theory of evolution. The story of the Meselson–Stahl experiment, as told here by its protagonists, also reveals how friendship and overcoming obstacles are as crucial to the scientific process as ideas themselves.
Learning Overview —
Big Concepts
Faithful replication of the genetic material (DNA) is the foundation of all life on earth. The experiment by Meselson and Stahl established that DNA replicates through a semi-conservative mechanism, as predicted by Watson and Crick, in which each strand of the double helix acts as a template for a new strand with which it remains associated, until the next replication.
Bio-Dictionary Terms Used
Bacteriophage (phage), base, base pairing, chromosome, DNA, Hershey–Chase experiment, eukaryote, mutation, nucleotides, Prokaryote (bacteria), recombination, RNA, ultraviolet light
Terms and Concepts Explained
Equilibrium density-gradient centrifugation, DNA replication, isotope, semi-conservative DNA replication
Introduction
Matthew Meselson and Franklin Stahl (both 24 years old) met at the Marine Biological Laboratory in Woods Hole in Massachusetts and decided to test the Watson–Crick model for DNA replication, which was unproven at the time.
What Events Preceded the Experiment?
Watson and Crick proposed a "Semi-Conservative" model for DNA replication in 1953, which derived from their model of the DNA double helix. In this proposal, the strands of the duplex separate and each strand serves as a template for the synthesis of a new complementary strand. Watson's and Crick's idea for DNA replication was a model, and they did not have data to support it. Some prominent scientists had doubts.
Two other models, "Conservative" and "Dispersive", for DNA replication were proposed.
Setting Up the Experiment
A method was needed to detect a difference between the parental and daughter (newly replicated) DNA strands. Then, one could follow the parent DNA molecule in the progeny. Meselson thought to distinguish between parental and newly synthesized DNA using a density difference in the building blocks (nucleotides) used to construct the DNA. The three models for DNA replication would predict different outcomes for the density of the replicated DNA in the first- and second-generation daughter cells.
The general experimental idea was first to grow bacteria in a chemical medium to make high-density DNA and then abruptly shift the bacteria to a low-density medium so that the bacteria would now synthesize lower density DNA during upcoming rounds of replication. The old and newly synthesized DNA would be distinguished by their density.
To measure a density difference in the DNA, Meselson and Stahl invented a method called equilibrium density gradient centrifugation. In this method, the DNA is centrifuged in a tube with a solution of cesium chloride. When centrifuged, the cesium chloride, being denser than water, forms a density gradient, reaching a stable equilibrium after a few hours. The DNA migrates to a point in the gradient where its density matches the density of the CsCl solution. Heavy and light DNA would come to different resting points and thus physically separated.
Doing the Key Experiment
Meselson and Stahl first decided to study the replication of DNA from a bacteriophage, a virus that replicates inside of bacteria, and used a density difference between two forms of the nucleobase thymine (normal thymine and 5-bromouracil). These experiments did not work.
The investigators changed their plans. They studied replication of the bacterial genome and used two isotopes of nitrogen (15N (heavy) and 14N (light)) to mark the parental and newly synthesized DNA.
When the population of bacteria doubled, Meselson and Stahl noted that the DNA was of an intermediate density, half-way between the dense and light DNA in the gradient. After two doublings, half of the DNA was fully light and the other half was of intermediate density. These results were predicted by the Semi-Conservative Model and are inconsistent with the Conservative and Dispersive Models.
Meselson and Stahl did another experiment in which they used heat to separate the two strands of the daughter DNA after one round of replication. They found that one strand was all heavy DNA and the other all light. This result was consistent with the Semi-Conservative model and provided additional evidence against the Dispersive Model.
Overall, the results provided proof of Semi-Conservative replication, consistent with the model proposed by Watson and Crick.
What Happened Next?
Within a couple of weeks after their key experiment, Meselson wrote a letter to Jim Watson to share news of their result (letter included).
Max Delbruck, the Caltech physicist and biologist who had proposed the dispersive model, was elated by the results, even though Meselson and Stahl disproved his replication hypothesis, and urged the young scientists to write up their results for publication and announce the important result to the world (1958).
Scientists now know a great deal about the protein machinery responsible for DNA replication.
Closing Thoughts
The Meselson–Stahl experiment had a powerful psychological effect on the field of genetics and molecular biology. It was the first experimental test of the Watson and Crick model, and the results clearly showed that DNA was behaving in cells exactly as Watson and Crick predicted.
In addition to having a good idea, the behind-the-scenes tour of the Meselson–Stahl experiment reveals that friendship and persistence in overcoming initial failures play important roles in the scientific discovery process. Also important was an atmosphere of freedom that allowed Meselson and Stahl, then very junior, to pursue their own ideas.
Guided Paper
Meselson, M. and Stahl, F.W. (1958). The replication of DNA in Escherichia coli. Proceedings of the National Academy of Sciences U.S.A., 44: 672–682.
This classic paper provides experimental evidence that the strands of the DNA double helix serve as templates to create a new copy of DNA. These results provide experimental evidence of The Watson and Crick model of DNA replication (‘semi-conservative replication’) demonstrating that genetic information is passed from one cell or organism to its progeny.
DownloadIntroduction
The first conversation between Matt Meselson and Frank Stahl, in the summer of 1954, began a collaboration that led to their Key Experiment on DNA replication and marked the beginning of a lifelong friendship. Matt and Frank describe the circumstances that brought them together below.
Matt
It was 1954, the year after Jim Watson and Francis Crick published their two great papers describing their double helical model of DNA and its implications for how it might replicate, mutate, and carry genetic information. Jim Watson (26 years old) and I (a 24-year-old first-year graduate student) were both at Caltech and living at the Athenaeum, the Caltech faculty club. We often talked while waiting for dinner. One day, Jim asked me to join him for the summer as his teaching assistant in the Physiology Course at the Marine Biological Laboratory (MBL) at Woods Hole, Massachusetts. He mainly wanted me to do experiments to see if RNA was a double helix. (As a side note, those experiments showed that RNA is not a double helix.) So in June 1954, I drove my 1941 Chevrolet coupe across the country from Cal Tech in Pasadena, California, to the MBL at Woods Hole, Massachusetts.
One day in Woods Hole, while planning student assignments for the Physiology Course, Jim went to the window of the course office upstairs in the MBL Lillie building and pointed towards a student sitting on the grass under a tree serving gin and tonics. That student was Frank Stahl. Jim said let's give him a really hard experiment to do all by himself – the Hershey–Chase Experiment; then, we'll see how good he really is. Two years earlier, Alfred Hershey and Martha Chase published an influential experiment that provided evidence implicating DNA, and not protein, as the substance conferring genetic inheritance in bacteriophage (see the whiteboard animation video on the Hershey–Chase experiment).
I thought that this guy serving gin and tonics must be an interesting fellow, so I went downstairs to meet him and let him know what was being planned for him. Frank was then a biology graduate student at the University of Rochester. I sat down on the grass under the tree, and we hit it off right away. We found that we had much to discuss. I was very impressed with Frank's knowledge of phage genetics, a subject I knew nothing about, coming from the laboratory of Linus Pauling where I was doing X-ray crystallography. Frank can tell you more about our conversation under the "gin and tonic tree."
Frank
1954 was an exciting time for molecular biology. One year earlier, Watson and Crick published their model for the double helix structure of DNA (see the Narrative on DNA Structure by Vale), which aroused much excitement as well as some serious disbelief. With Watson as an instructor and Crick and Sydney Brenner as visitors and several other founders of molecular biology, Woods Hole that summer became an epicenter for discussion of the great questions in molecular biology. Could the double helix model, as Watson and Crick proposed, explain the replication of the genetic information? What is the "code" for reading out the nucleotide sequence of DNA and turning that into the sequence of amino acids in protein? Would RNA have a similar structure to DNA? However, at the time of my arrival, I had no idea that Woods Hole was hosting anyone, but me, interested in the big problems of modern, i.e., molecular, biology.
I was a 24-year-old biology graduate student at the University of Rochester and had come to the MBL to take the Physiology Course. To beat the heat and, perchance, to meet someone interesting, I invested in a bottle of gin and a 6-pack of tonic, found some ice, a thermos jug, and a tree and sat myself down in the shade. Matt Meselson was one of my first catches.
Our conversation under the "gin and tonic tree" was life-changing. After Matt warned me of Watson's planned test of my experimental aptitude, we talked about the work we were doing. I explained to him the problem in phage genetics on which I was working. Eventually our conversation turned to DNA replication. Neither of us was working on the problem at the time, although we were both keenly interested in it. It was perhaps the most important contemporary question in molecular biology.
At Caltech, Matt had already come up with an idea for how the mechanism of DNA replication might be studied by density labeling. But, as a physical chemist, he was unfamiliar with the methods of phage and bacterial biology that would be needed to conduct the actual experiments. So we decided to collaborate. I was planning to be a postdoctoral fellow at Caltech starting that September. If we could develop a method for measuring the density of DNA molecules and successfully apply it to the problem of DNA replication, we could establish whether the Watson–Crick model for DNA replication, and even the model of the structure itself, was correct or not.
We did not begin our collaboration immediately because Matt needed to finish his X-ray crystallography and I had made plans to do my postdoctoral work on bacterial genetics with Joe Bertani.
What Events Preceded the Experiment?
You can also hear Matt Meselson describing the Meselson–Stahl experiment in Video 1.
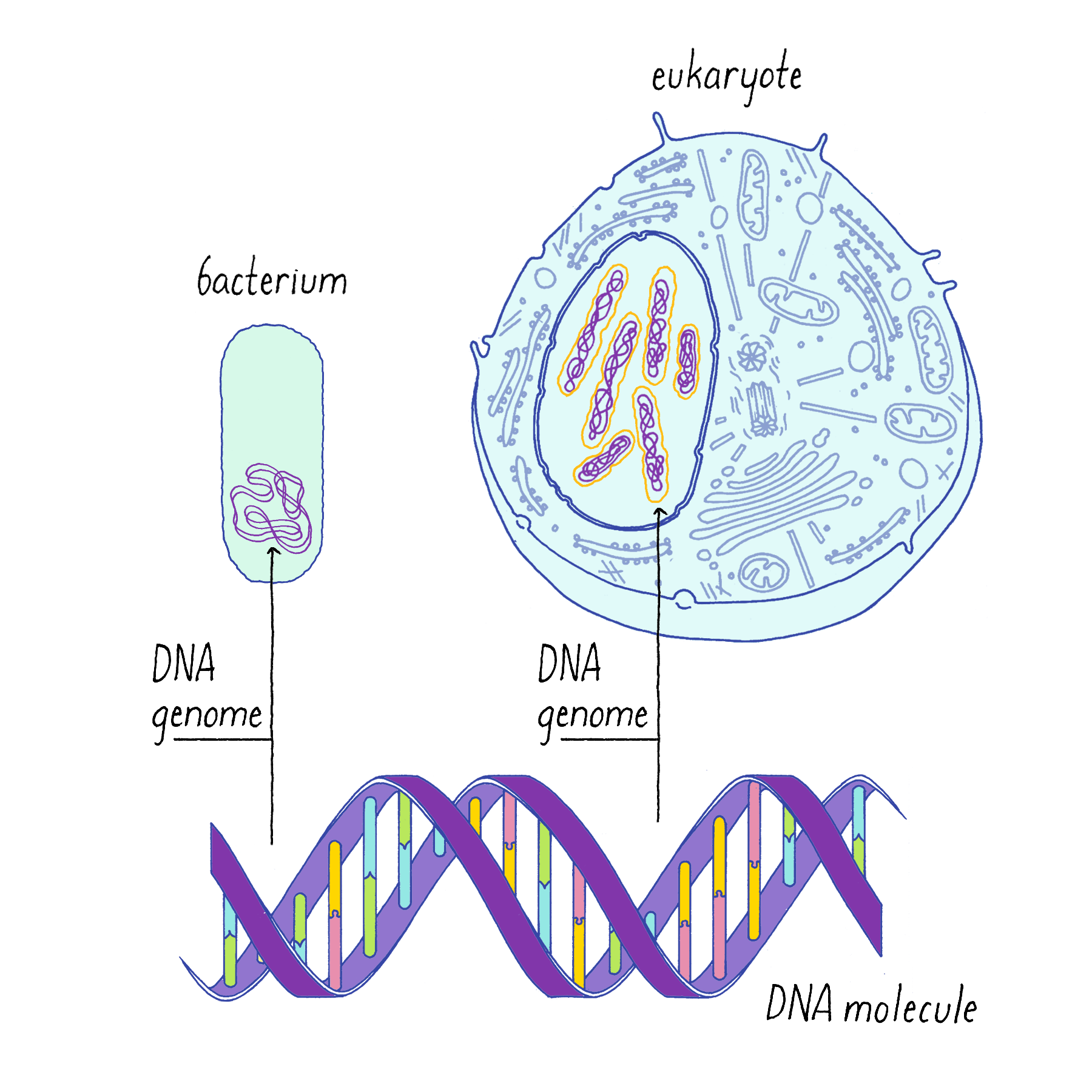
The genetic material of eukaryotic cells is organized in the form of chromosomes, each a single linear, double-stranded DNA molecule (Figure 1). Most prokaryotes (bacteria) have a single, circular chromosome (Figure 1). All forms of life must replicate their DNA and, except for recombination and infrequent mutations, pass identical copies of their genetic material to their progeny. (See also the Whiteboard Video on Keeping Track of Your DNA.)
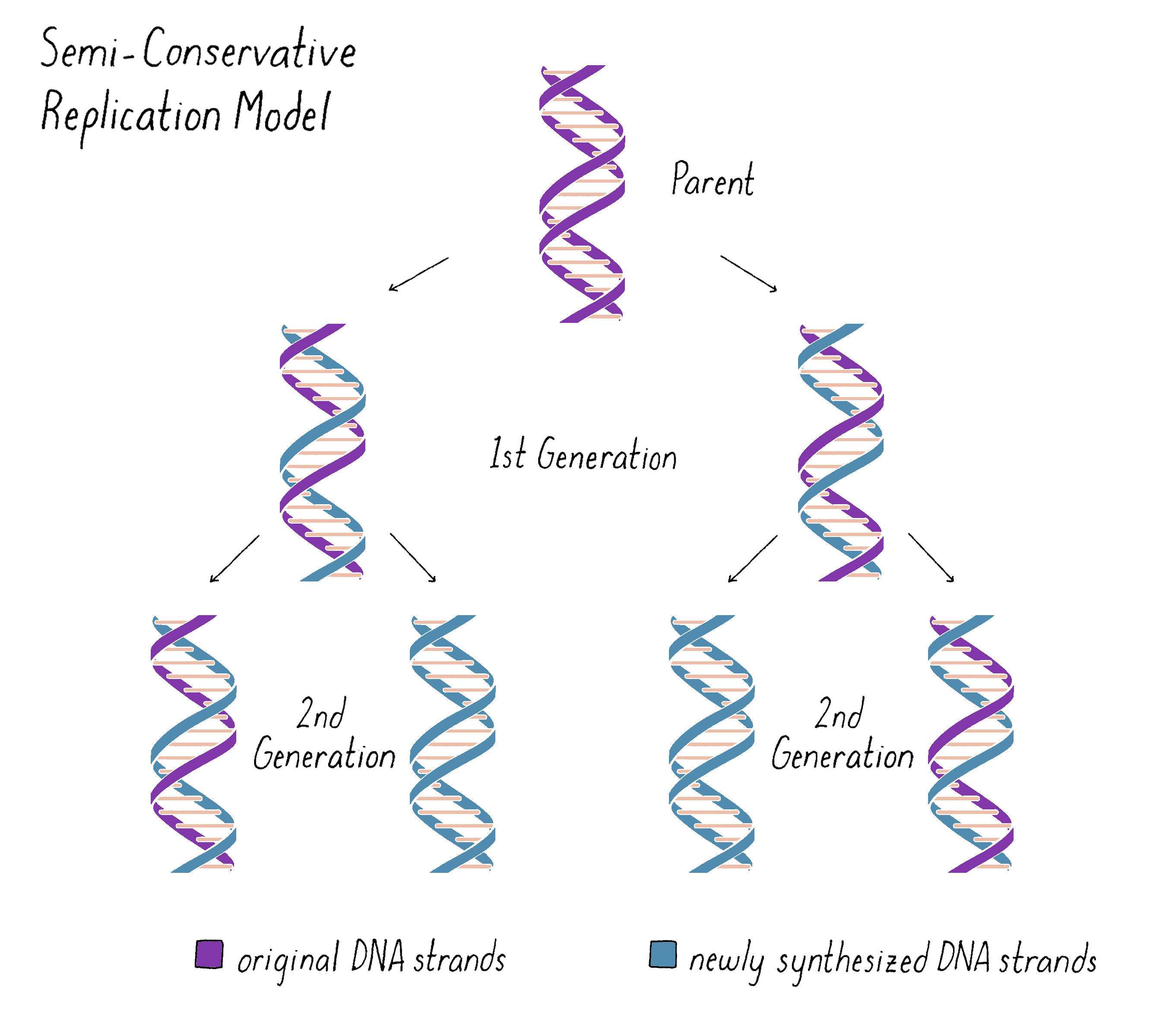
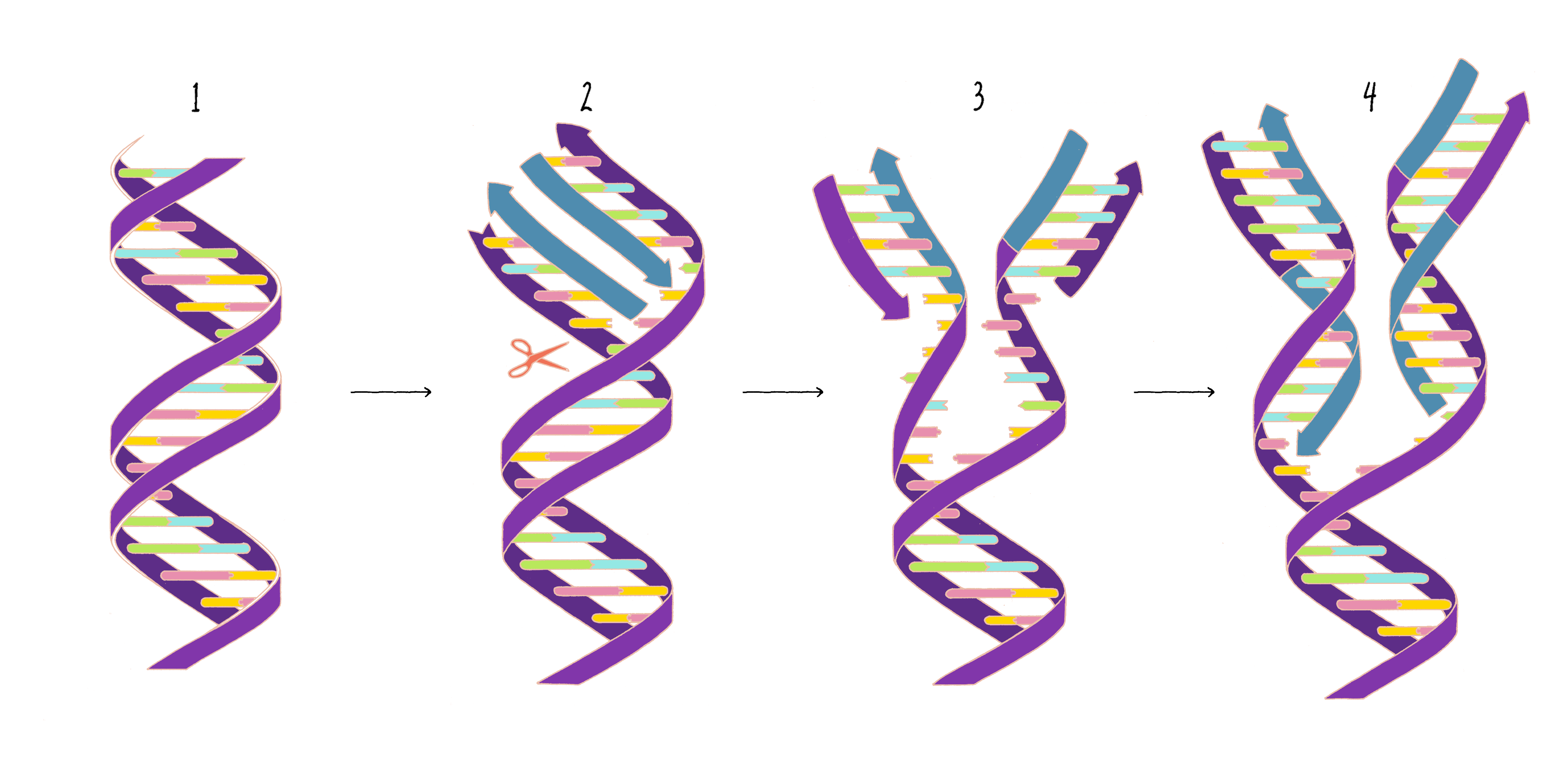
Based upon their model for the structure of DNA, Watson and Crick proposed that DNA replicates in a "Semi-Conservative" manner (Figure 2). In this model, the two strands of the DNA double helix unwind and separate, and each "parent" strand serves as a template for the synthesis of a new "daughter" strand. The Watson–Crick base pairing (see the Narrative on DNA Structure by Vale) of adenine with thymine and guanine with cytosine ensures that, except for rare copying errors, mutations, the information of the original double-stranded DNA molecule is preserved during the synthesis of the daughter strands. In the Semi-Conservative model, each daughter cell in the first generation would inherit one of the original DNA strands from the parent and a recently synthesized DNA strand. In the second generation, two of the granddaughters would be composed of all newly synthesized DNA and two granddaughters would have hybrid DNA (one parental strand and one newly synthesized strand).
While (spoiler alert) the Semi-Conservative Model turned out to be correct, it was far from a foregone conclusion. Before our experiment, several leading scientists questioned the Semi-Conservative Model (see Dig Deeper 1 for more information on their reservations) and proposed alternate models discussed below.
Explorer's Question: What do you imagine are the pros and cons of this model?
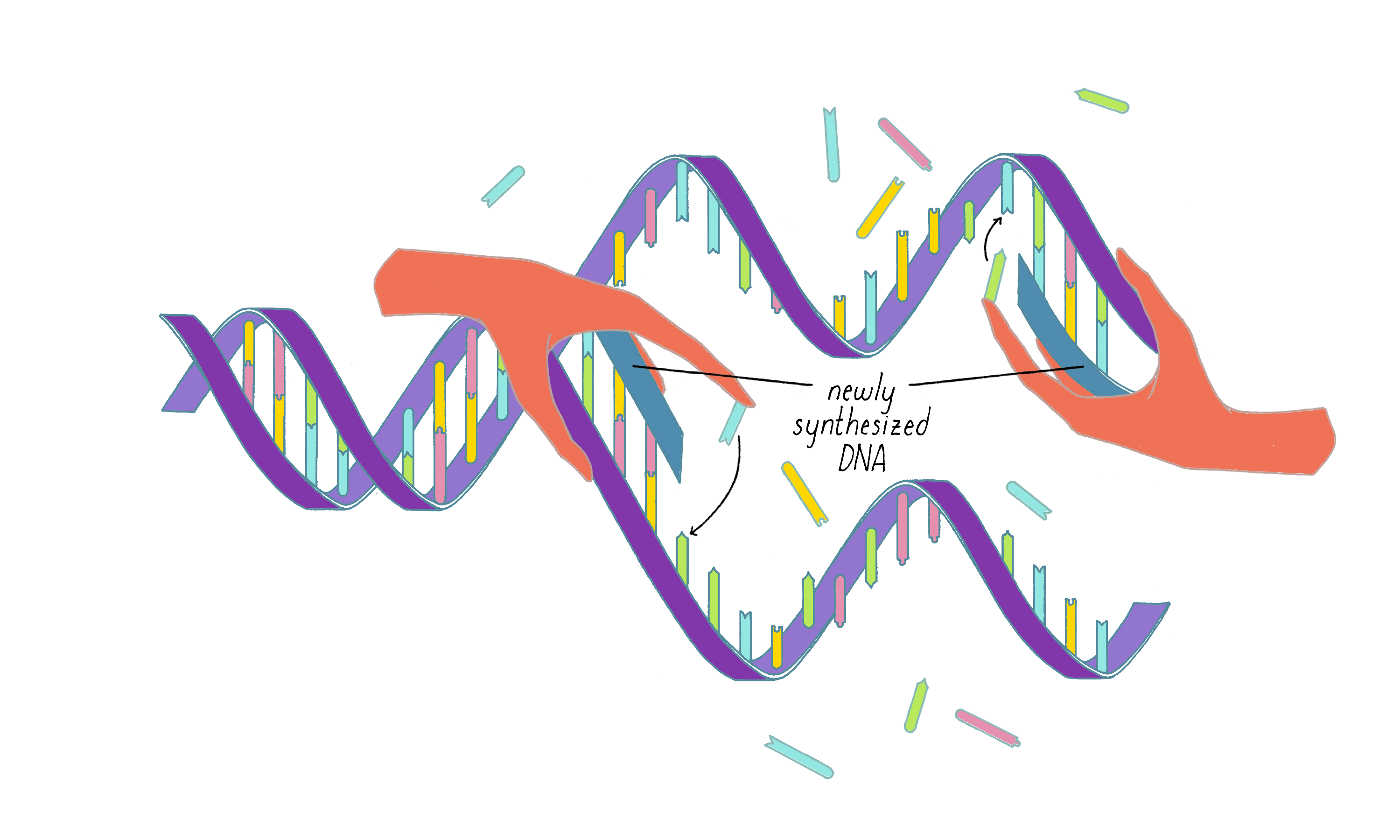
Answer: The beauty of this model is that it provides a clear explanation of how a daughter strand is made from the template strand (Figure 3). However, the model requires that the two long parental DNA strands unwind to become single-strand templates. This was seen as a weakness by many scientists at the time (see Dig Deeper 1 for more information).
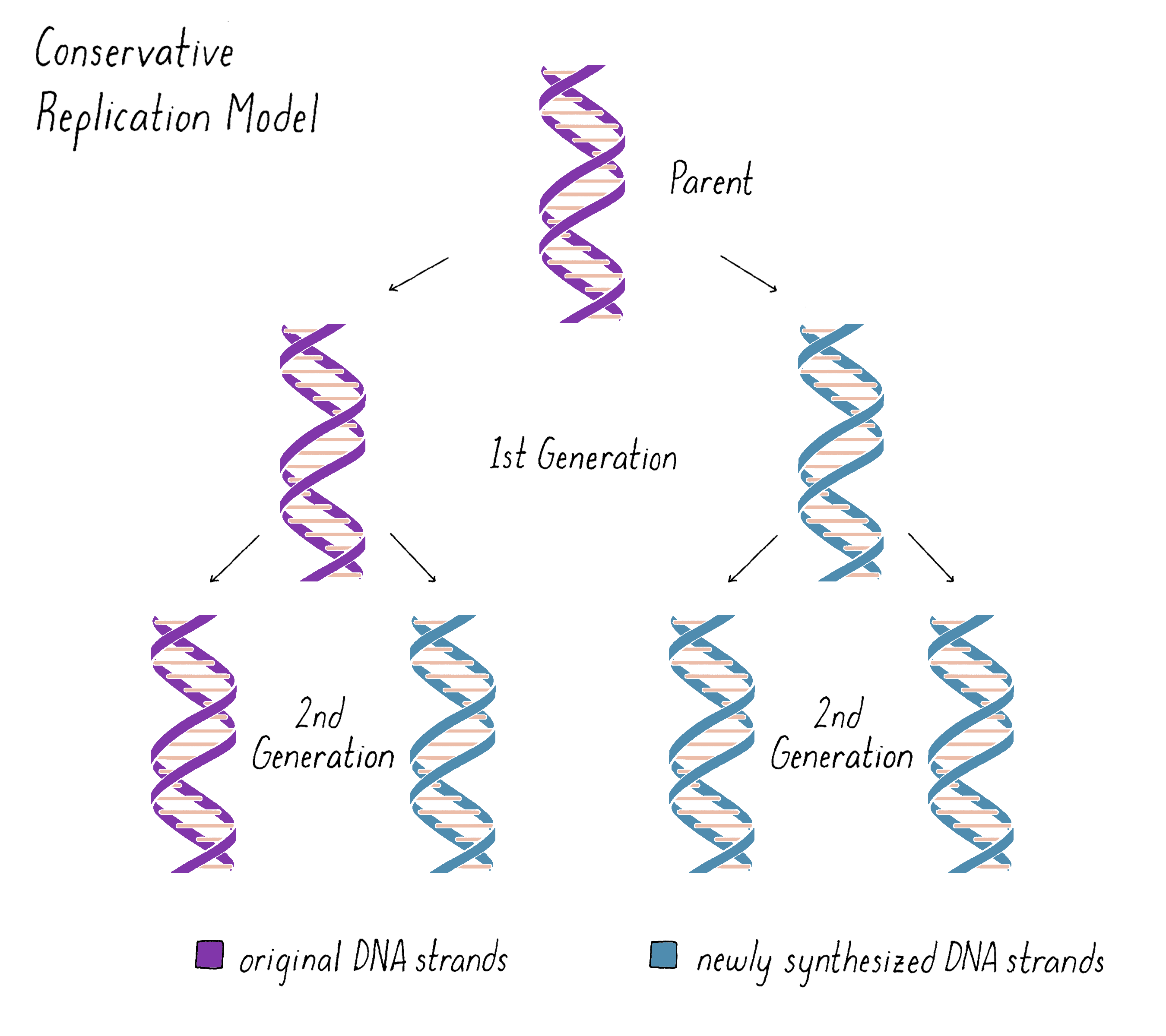
The unwinding of the DNA helix and keeping the daughter and parental strands from becoming tangled posed problems for the Semi-Conservative model in the minds of many scientists. As a result, other models for DNA replication were imagined. One was a "Conservative Replication" model (Figure 4). In this model, the parental double helix forms a template for a completely new double helix. The two original strands remain together, no unwinding occurs, and the daughter DNA is formed from newly synthesized DNA. In this model, in the first generation, one daughter DNA would inherit the original DNA double helix from the parent DNA and the other daughter DNA would inherit the newly synthesized DNA double helix. In the second generation, one of the four granddaughters would have the original parental DNA and the other three granddaughters would all be composed of newly synthesized DNA. While Conservative Replication was a logical possibility, it was not elaborated by any specific mechanism.
Explorer's Question: What do you imagine are the pros and cons of this model?
Answer: This model did not call for unwinding of the DNA strands, as in the Semi-Conservative Model, thus solving the concern about DNA unwinding. However, it was unclear how the copying machinery would read out the nucleotide sequence information buried in the core of the DNA double helix.
A third possibility was a model proposed by Max Delbrück (later called "Dispersive Replication") (Figure 5). Delbrück doubted that the two strands of the double helix could be unwound or pulled apart to undergo Semi-Conservative replication and instead suggested that DNA strands broke at every half-turn of the helix during replication (discussed in more depth below and in Dig Deeper 1). According to Delbrück's Dispersive Replication Model, each helix of the replicated DNA consists of alternating parental and daughter DNA. Unlike the other two models, the progeny in subsequent generations would be indistinguishable with regard to their compositions of parental and newly synthesized DNA.
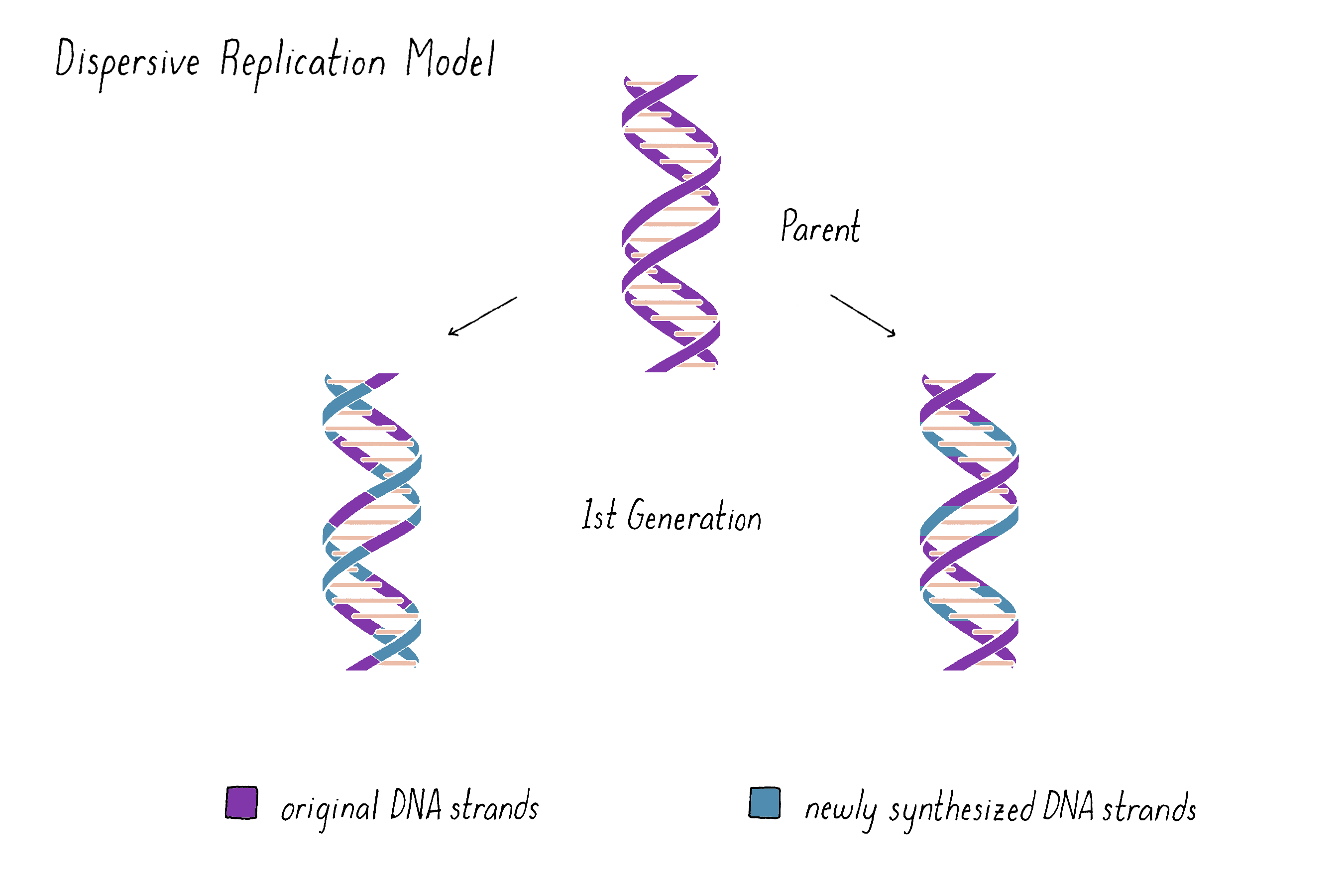
Explorer's Question: What do you imagine are the pros and cons of this model?
Answer: Fragmentation would create shorter templates for replication, which would minimize any unwinding or untangling problems faced by one very long DNA molecule. However, the reassembly of the fragments again into the intact chromosome could be problematic, especially if the breaks occur at every half-turn of the helix.
Explorer's Question: In the first generation, which model(s) would predict that the two daughter cells would receive approximately equal amounts of the original and newly synthesized DNA?
Answer: The Semi-Conservative Model and the Dispersive Model. However, differences in daughter composition arise in the second generation in the two models.
Matt and Frank learned about the models for DNA replication prior to their first meeting at the Physiology Course at the Marine Biological Laboratory through circumstances described below.
Matt
Sometime in 1953, while I was a graduate student of the great chemist Linus Pauling, I went to see Max Delbrück, a physicist and founder of the "phage group" who had become deeply interested in genetics and the basis of life (Max Delbrück, and his work with Salvador Luria, is featured in the Narrative on Mutations by Koshland). I wanted to learn what problems in biology he thought were most important and what advice he might have for me about getting into biology. Almost as soon as I sat down in his office, he asked what I thought about the two papers by Watson and Crick that had been published in Nature earlier that year. I confessed that I had never heard of them.
Exasperated, Delbrück flung a little heap of reprints of the Watson–Crick papers at me and shouted "Get out and don't come back until you have read them." What I heard was "come back." So I did, but only after reading the papers.
When I came back, Delbrück said he did not believe in the Semi-Conservative mechanism of DNA replication proposed by Watson and Crick. Max had imagined that if replication is semi-conservative, the two daughter duplexes would become wound around each other turn-for-turn as the two chains of the mother molecule became unwound. To get around the supposed problem of untangling the daughter molecules, he proposed a model in which breaks are made to prevent interlocking when separating, and then joined back together (see Figure DD1 in Dig Deeper 1). This mechanism required rotation of only short lengths of duplex DNA, after which the chains would be rejoined. In the rejoining process, sections of the new chain would be fused to sections of the old chain, making all four of the chains mosaics of new and old DNA. Delbrück, in 1954, published a paper that questioned the Watson–Crick model and presented this new model (later referred to as "Dispersive Replication" as shown in Figure 5 and Dig Deeper 1). In some ways, the idea of Delbrück was ahead of its time. Transient breaks are now known to be made by an enzyme called topoisomerase, but in a manner that leaves the individual chains of the parent duplex intact (see Dig Deeper 1).
In addition to Max's reservations and model, several other scientists also posed their own concerns and solutions to the "unwinding problem" or alternatives to the Watson–Crick DNA structure itself (see Dig Deeper 1).
What I gathered from my conversations with Max and others was that not everyone believed the DNA replication model of Watson and Crick. It was only a hypothesis with no experimental evidence to support it. The key to solve this problem was to follow the parental DNA in the progeny. But how?
I was working on something very different for my PhD thesis with Linus Pauling, but earlier that year, I had an idea for labeling protein molecules with deuterium, a heavy isotope (2H) of hydrogen (1H) and separating them from unlabeled protein molecules in a centrifuge according to their density as a means to solve a quite different problem (see Dig Deeper 2). After that second meeting with Max, it occurred to me that density labeling and centrifugal separation might be used to solve the DNA replication problem. When I told this to Pauling, he urged me to get my X-ray crystallography done first. And when I proposed the density approach to Watson, one of those evenings waiting for dinner at the Athenaeum, he said I should go to Sweden to do it – where the ultracentrifuge had been invented (which I never did).
Frank
My entry point to thinking about DNA replication came when I was trying to understand how bacteriophage (viruses that infect bacteria) exchange pieces of DNA with one another. This process of DNA exchange between chromosomes is called recombination (see the whiteboard video on the experiments by Morgan and Sturtevant). When did this recombination process occur? Did it occur when DNA replicates? Or perhaps recombination was an event that stimulated DNA replication? My intuition was that the processes of recombination and replication were somehow related. However, little was known about the mechanisms of either DNA replication or recombination at the time. Furthermore, I did not know how to pursue these questions in 1954. My ideas for experiments were lame and would have led to un-interpretable data.
Like many interesting questions in biology, often one has to be patient until either the right idea or technology emerges that allows one to answer them properly. In 1954, my awareness of a possible connection between replication and recombination primed my interest for the first gin and tonic conversation with Matt. However, it was several decades before I was sure that, in bacteriophage, DNA replication and recombination, in a large degree, depended upon each other. The convincing experiments were based on variations of a technique pioneered by Matt and Jean Weigle at Caltech. In this method, density-labeled, genetically marked parental phage infect the same bacteria. The densities and genetic makeup of progeny phage are determined by bioassay of the individual drops collected from a density gradient.
Setting Up the Experiment
Matt and Frank
To distinguish between the Semi-Conservative, Conservative, and Dispersive Models of DNA replication described above, we needed a method that could tell the difference between the parental and daughter DNA strands. Figures 2, 4, and 5 illustrate the parent and newly synthesized strands with different colors. However, we needed to find a real physical difference that would serve the same function of distinguishing between the old and newly synthesized DNA. Matt had the idea of distinguishing old and new DNA by having the bacteria synthesize them with different isotopes and separating them in a centrifuge according to their density. If the original and the newly synthesized DNAs could be made of different density materials, then we could perhaps measure this physical difference. We will discuss the chemicals that were used to make the DNA heavier or lighter in the next section.
Our experimental idea was to grow an organism in a chemical medium that would make its DNA heavy. Then, while it was growing, we would switch to a new medium in which the newly synthesized DNA would be made of lighter material (Figure 6). The density difference between the original and the newly replicated DNA could allow us to distinguish between models for DNA replication.
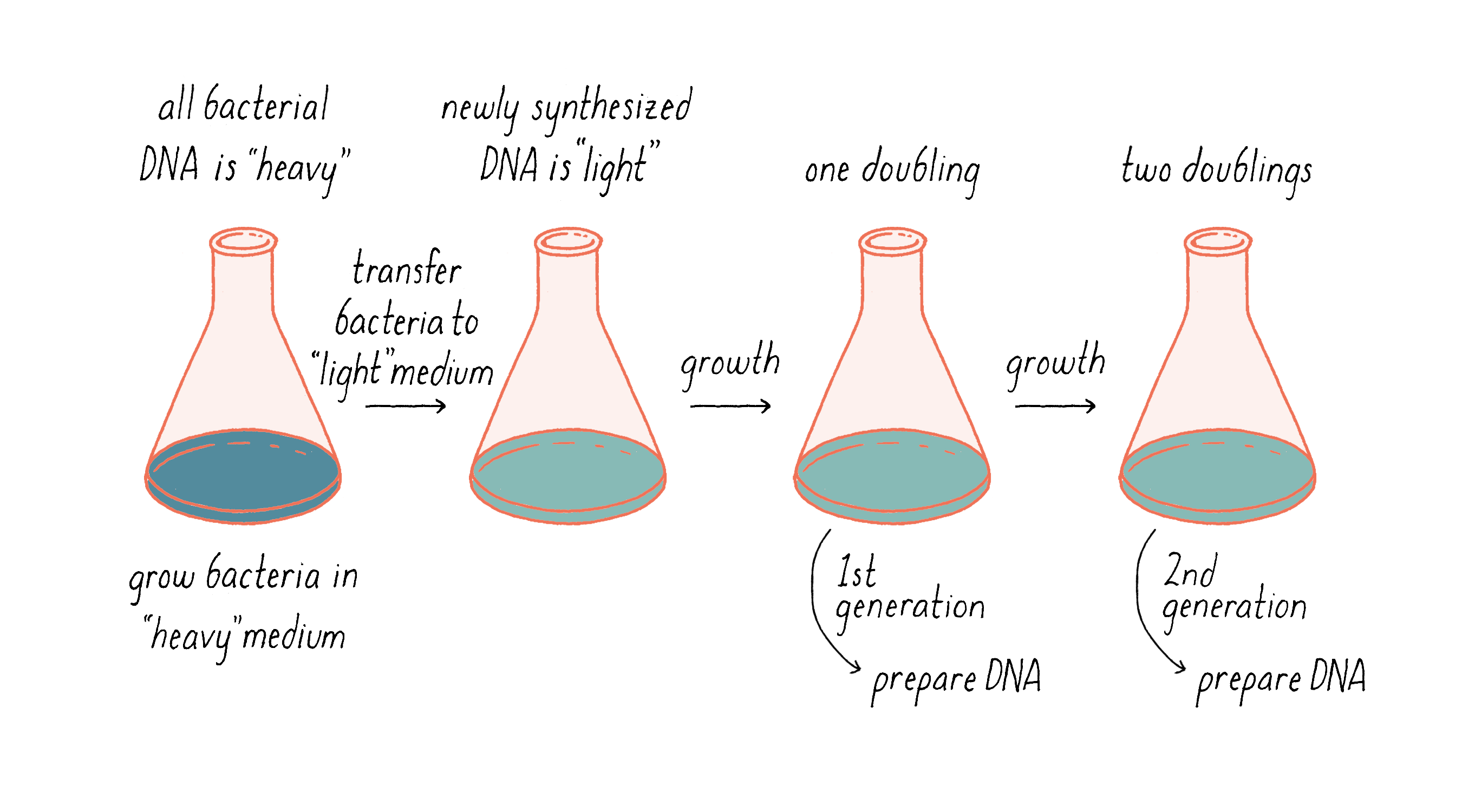
To separate DNA of different density, we invented a method, called "equilibrium density gradient centrifugation," and published it, together with Jerome Vinograd, a Caltech Senior Research Fellow who had taught us how to use the ultracentrifuge in his lab and provided advice. In this method, as applied to DNA, a special tube that has quartz windows so that ultraviolet light photos can be taken while the centrifuge is running is filled with a solution of cesium chloride and the DNA to be examined. Upon centrifugation at high speed (~45,000 revolutions per minute or 140,000 times gravity), the CsCl gradually forms a density gradient, becoming most concentrated at the bottom of the tube (Figure 7). The CsCl solution toward the top of the tube is less dense than the DNA, while the CsCl solution at the bottom is denser than DNA. Thus, when a mixture of DNA in a CsCl solution is centrifuged, the DNA will eventually come to a resting point where its density matches that of the CsCl solution (Figure 7). The DNA absorbs UV light, and its position along the tube was recorded by using a special camera while the centrifuge is running.
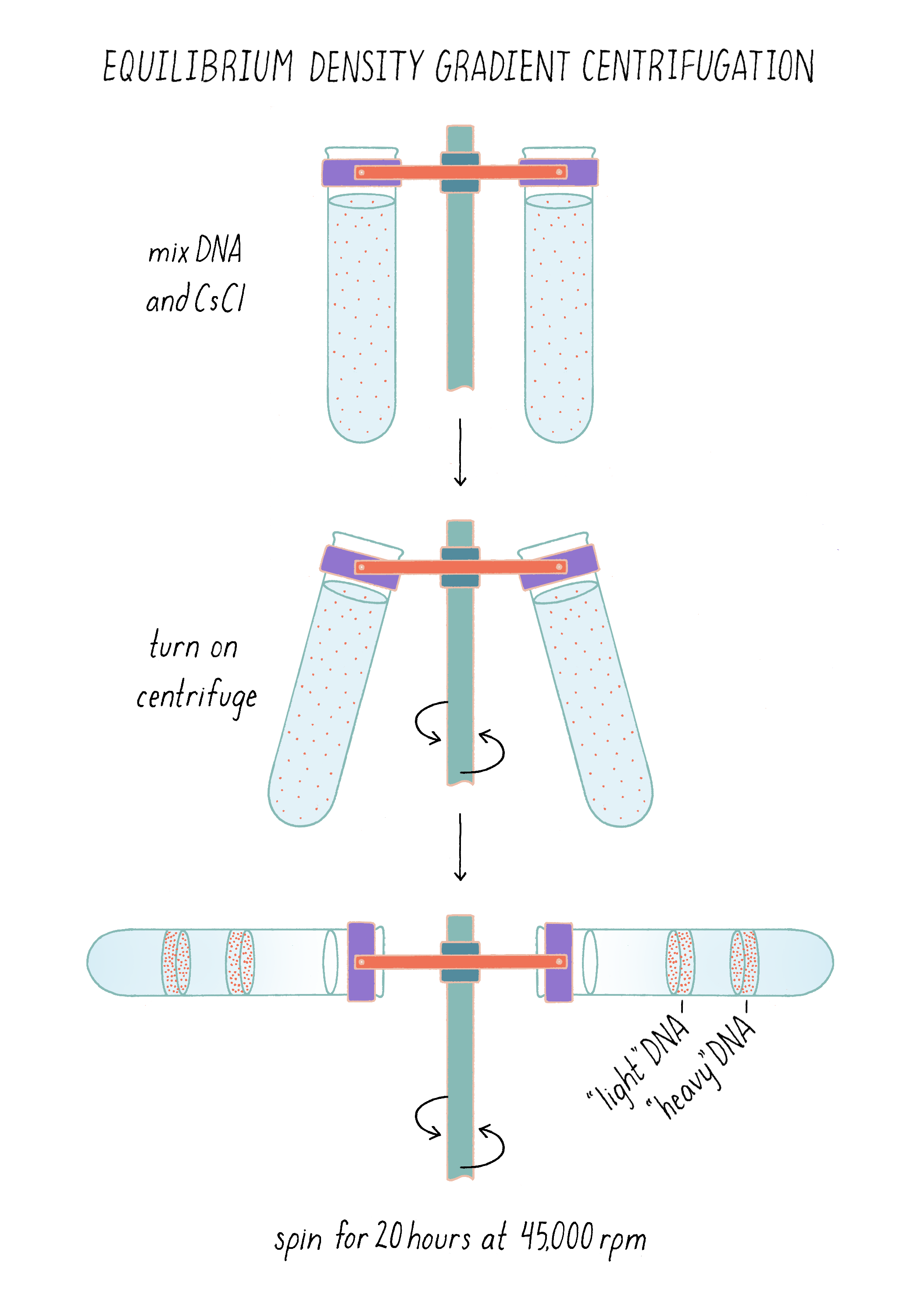
The method now seems straightforward, but in reality, it took a couple of years to develop. For example, we did not come to cesium chloride immediately. We looked at a periodic table for a dense monovalent atom that would not react with DNA; rubidium chloride (molecular weight of 121) was available in the Chemistry Department stockroom and we initially tried to use that but found that even concentrated solutions were not dense enough to float DNA to a banding point. We then moved one level down in the periodic table to cesium (the molecular weight of cesium chloride is 168) and that worked (for more details, see Dig Deeper 3).
Doing the Key Experiment
Frank and Matt
In the fall of 1954, we were reunited in Cal Tech and lived for about eight months in the same house across the street from the lab. We finally could begin doing experiments to test models of DNA replication. It should be noted that DNA replication was our "side" project; we also had our "main" projects under the supervision of our respective professors. However, faculty at Cal Tech was kind in allowing two young scientists to venture forward with their own ideas.
While the general experimental approach that took form under the "gin and tonic tree" was straightforward, choices had to be made in how exactly to do the experiment. What organism should we use? Would a chemical trick of making DNA heavier or lighter work and could we measure a small density difference between the two? It took us a while to get the conditions right, about two years.
We first decided to examine the replication of the bacteriophage T4 inside of the bacterium Escherichia coli. Bacteriophages are viruses that invade and replicate inside of a bacterium; when new viruses are made, they will burst the bacterium and then spread to new hosts. Bacteriophage have small genomes and are therefore the smallest replicating systems. Frank's PhD thesis was on T4, so he knew how to work with this phage. Max Delbrück and others at Cal Tech were also actively studying phage (see the Narrative on Mutations by Koshland). Thus, T4 seemed the obvious choice. To create DNA of heavier density than normal DNA, we decided to use the analogue, 5-bromouracil, of the base thymine, in which a heavier bromine atom replaces a lighter hydrogen atom. During replication, 5-bromouracil could be incorporated into DNA, instead of thymine.
However, while this approach seemed reasonable, it did not work in practice. Although we did not appreciate it at the time, during phage growth, the DNA molecules undergo recombination, joining parental DNA to newly synthesized DNA in a manner that after several generations would give no clear-cut distribution of old DNA among progeny molecules. Also, we learned from a recent paper that 5-bromouracil was mutagenic and made a detour into studying mutagenesis before coming back to our main project.
We clearly needed a new strategy.
Instead of phage, we decided to study the replication of the bacterial genome. This was a good choice – the bacterial DNA gave a very sharp and clear band when centrifuged in a solution of cesium chloride (to learn more about why we used cesium chloride to create a density gradient and the general use of this technique; see Dig Deeper 3).
We also switched our density label. DNA is made up of several elements – carbon, nitrogen, oxygen, phosphorus, and hydrogen. Some of these elements come in different stable isotopes, with atomic variations of molecular weight based upon different numbers of neutrons. Nitrogen-15 (15N) is a heavier isotope of nitrogen (the most common isotope, 14N, has a molecular weight of 14 Daltons). We could easily buy 15N in the form of ammonium chloride (15NH4Cl), which was the only source of nitrogen in our growth medium. The 15N in the medium then found its way into the bacterial DNA (as well as other molecules) in a harmless manner and did not impair bacterial growth.
We also had good luck in that Caltech bought a brand new type of ultracentrifuge called an analytical ultracentrifuge (Model E) developed by the Beckman Instrument Company. The Model E was a massive machine about the size of a small delivery truck (the current model is just a bit bigger than a dishwasher). Importantly, the Model E could shine a UV light beam on the tube while the centrifuge was spinning and detect and photograph the position of the DNA. The good news was that 15N-containing DNA and 14N-containing DNA could be clearly distinguished by their different density positions (Figure 8).
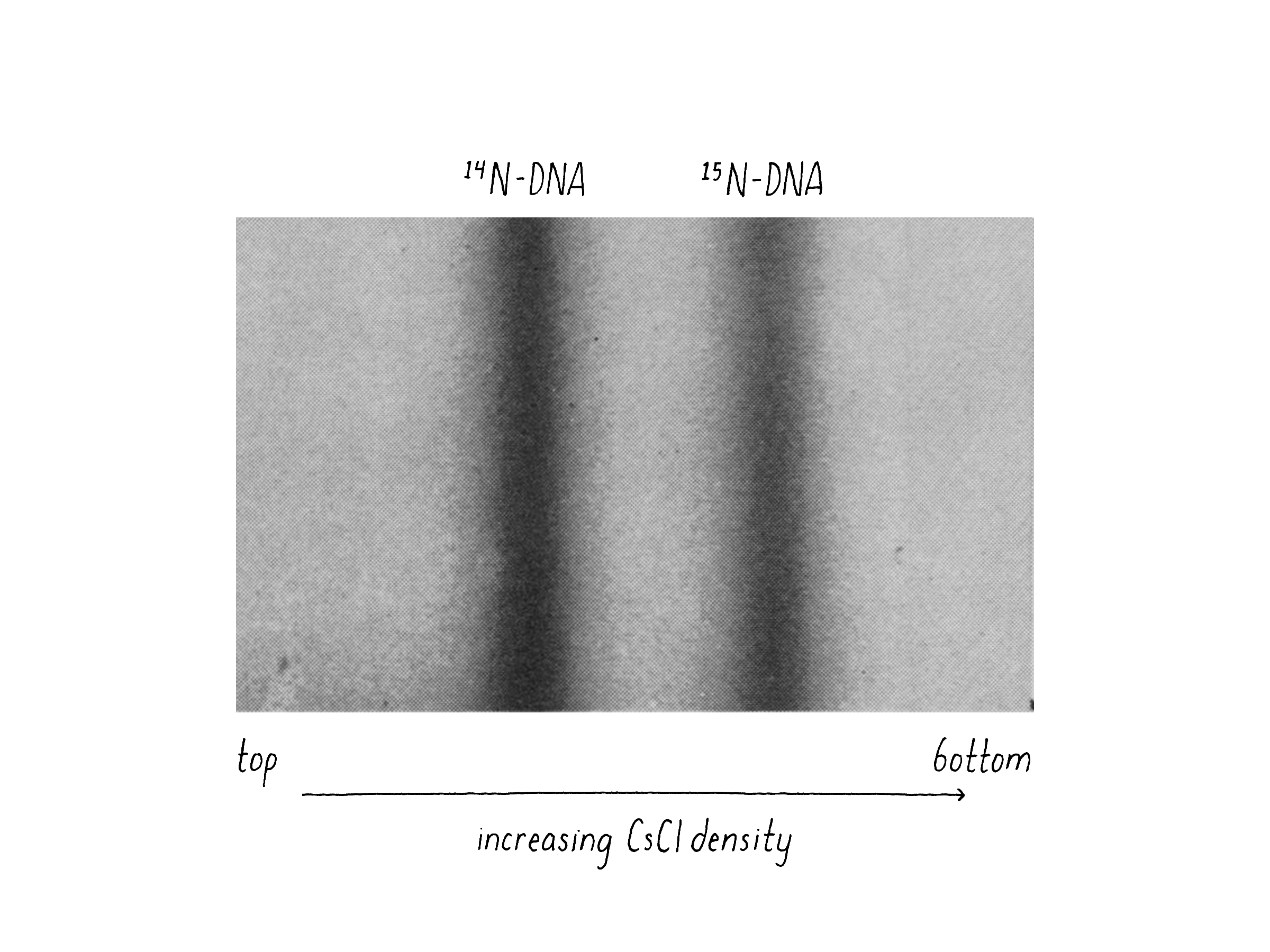
Matt
Finally, we had everything in place to try our experiment properly. I decided to set up our first experiment in the following two ways:
1) Grow the bacteria in "light" nitrogen medium and then switch to "heavy."
2) Grow another culture of bacteria in "heavy" nitrogen for many generations and then switch to "light."
Frank was called to a job interview and could not perform this first experiment with me. But before leaving, he warned me – "Don't do the experiment in such a complicated way on your first try. You might mix up the tubes."
I ignored Frank's advice and did the experiment both ways.
In the first experiment after transferring bacteria grown in heavy nitrogen (15N) growth medium and then switched to "light" (14N) nitrogen medium, I saw three discrete bands corresponding to old, hybrid, and new DNA, as predicted by Semi-Conservative replication. Excited developing the photograph in the darkroom, I remember letting out a yelp that caused a young woman working nearby to leave in a hurry. But later I realized my mistake. Frank had been prophetic. I indeed had mixed tubes, combining two different samples, one taken before and the other taken after the first generation of bacterial growth in the light medium. As described for the correct experiment below, there is no time when old, hybrid, and fully new DNA are present at the same time.
Frank
When I came back from my trip, Matt and I performed what proved to be the decisive experiment. We grew bacteria in "heavy" nitrogen (15N; from 15NH4Cl) and then switched to "light" nitrogen (14N; 14NH4Cl) and, at different time points, collected the bacteria by centrifugation, added detergent to release the DNA, and combined this with concentrated CsCl solution to reach the desired density. After 20 hours of centrifugation and the final density positions of the DNAs had come into view, we knew that we had a clean answer (Figure 9). The DNA from bacteria grown in heavy nitrogen formed a single band in the gradient. However, when the bacteria were shifted to a light nitrogen medium and then allowed to replicate their DNA and divide once (first generation), essentially all DNA had shifted to a new, "intermediate" density position in the gradient (Figure 9). This intermediate position was half-way between the all heavy and all light DNA. At longer times of incubation in light nitrogen, after the cells had divided a second time (second generation), a DNA band at lighter density was seen and there were equal amounts of the intermediate and light DNA.
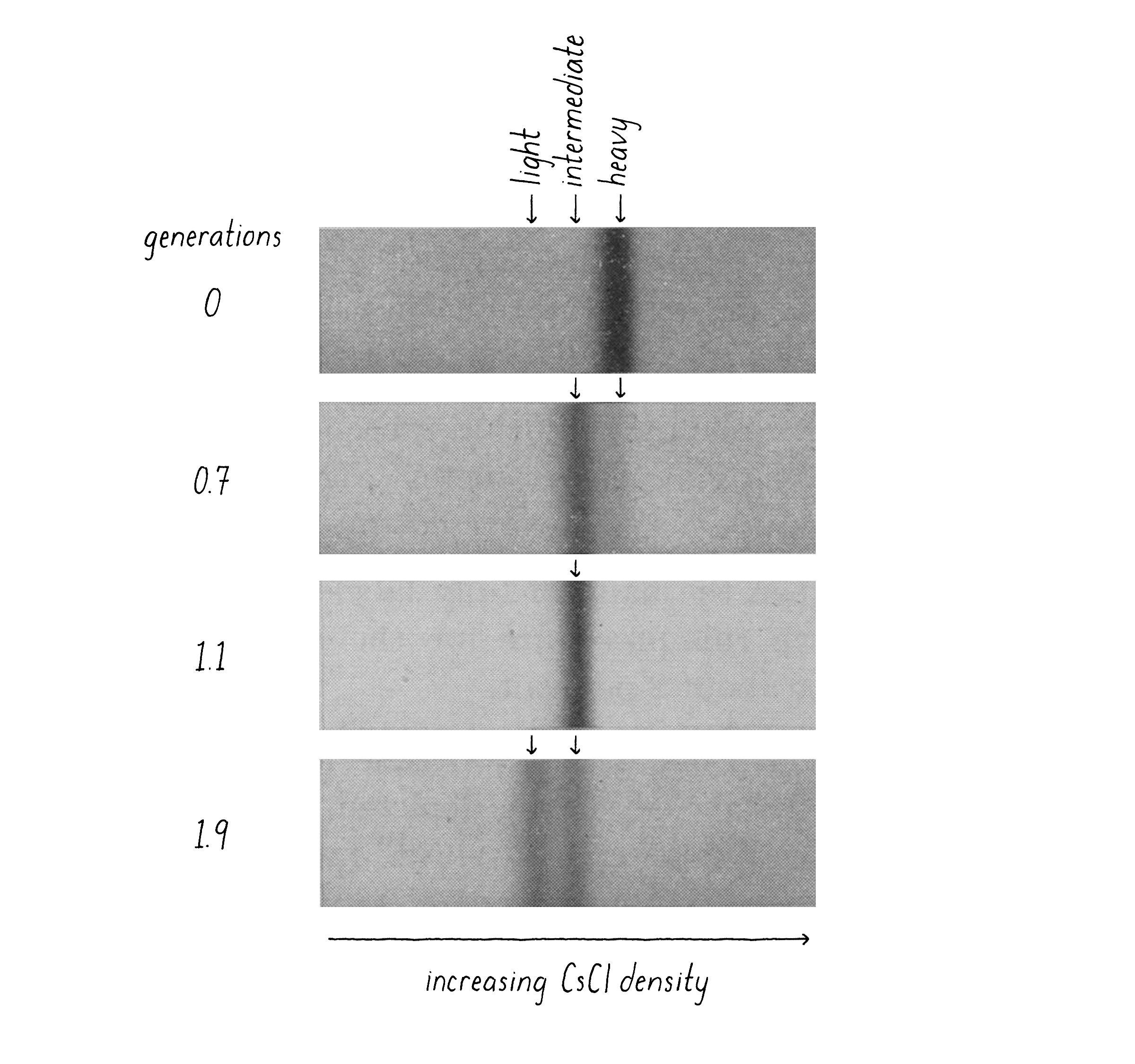
Explorer's Question: Which of the three models (Conservative, Semi-Conservative, or Dispersive) is most consistent with the results of this experiment?
Answer: The Semi-Conservative model. The Conservative model predicts a heavy and light band at the first generation, not an intermediate band. The Dispersive model predicts a single intermediate band at both the first and second generations (the band shifting toward lighter densities with more generation times).
Explorer's Question: Why are the two DNA bands at the 1.9 generation time point of approximately equal intensity?
Answer: After the first generation, each of the two heavy strands is partnered with a light strand. The bacterial DNA consists of one heavy strand and one light strand. When that heavy–light DNA replicates again in the light medium, the heavy strands are partnered with new light strands (intermediate density DNA) and the light strands are also partners with new light strands (creating all light density DNA).
Matt
The experiment that Frank described above took hardly any time at all (2 days) and yielded a clean result. We then repeated it without any problem. Once we knew how to set up the experiment, it was relatively easy. But it took us two years of trials before we got the experimental design and conditions right for the final ideal experiment.
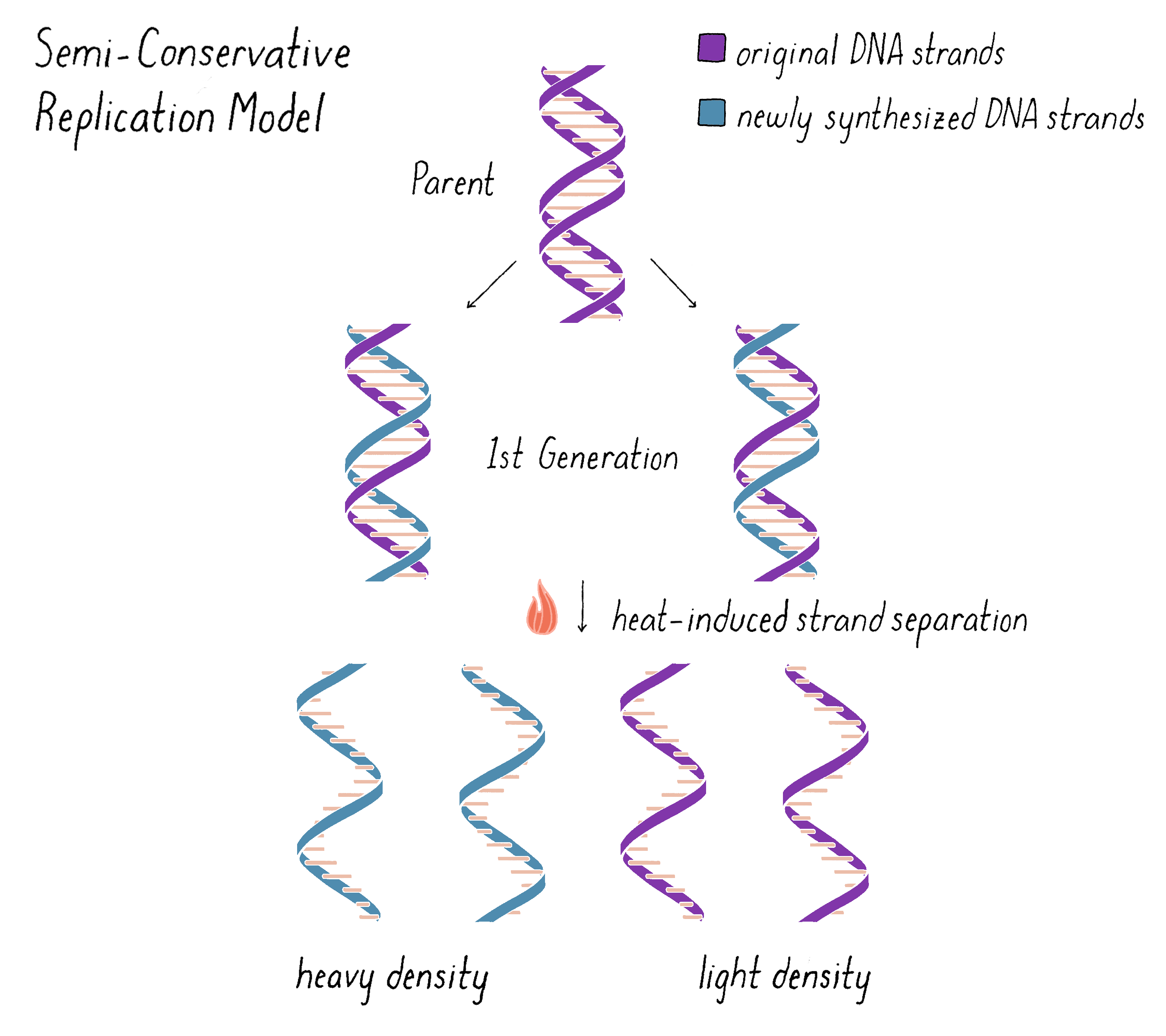
The experiment clearly supported the Semi-Conservative Replication model for replication and, in doing so, also supported the double helical model of DNA itself. However, we wanted to do one more experiment that would examine whether the "intermediate" density band of DNA in the first generation was truly made of two and just two distinct subunits, as predicted by the Watson–Crick model. The model predicts that one complete strand of DNA is from the parent and should be heavy and the other complete DNA strand should be all newly replicated and therefore light (Figure 10). We could test this hypothesis by separating the subunits with heat and then analyzing the density and molecular weight of the separated subunits by equilibrium density-gradient ultracentrifugation.
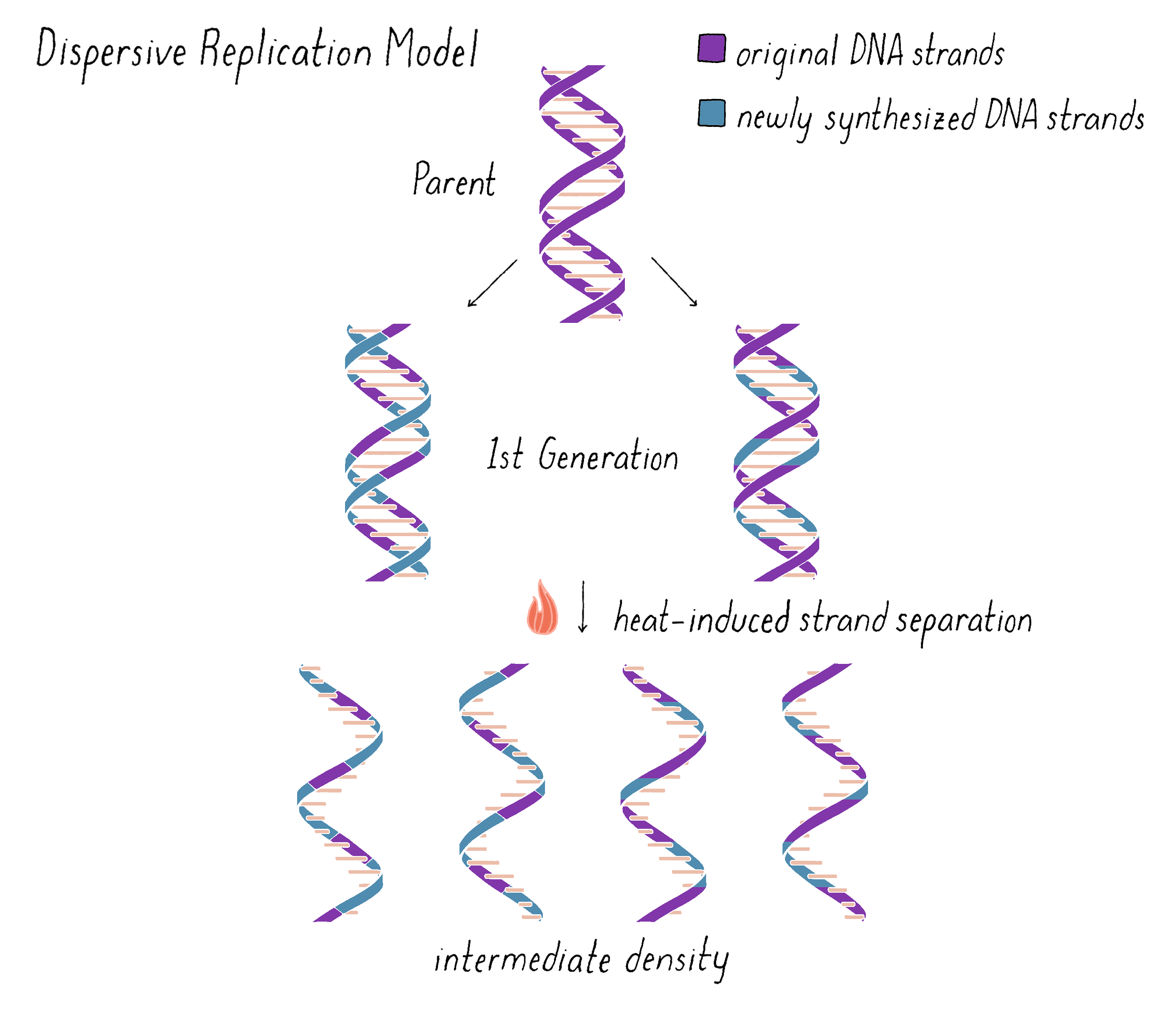
On the other hand, the Dispersive Model predicted that each DNA strand of first generation is an equal mixture of original and newly replicated DNAs (Figure 11).
The results from the experiment were again clear (Figure 12). The "intermediate density" DNA in the first generation split apart into a light and heavy component. From the width of the DNA band in the gradient (see Dig Deeper 3), we could also tell that the light and heavy DNA obtained after heating had each half of the molecular weight of the intermediate density DNA before heating. These results indicated that each parental strand remained intact during replication and produced a complete replica copy. This was decisive evidence against the Delbrück model for it predicted that both strands would be mosaics of heavy and light, not purely heavy and purely light. And the finding that the separated heavy and light subunits each had half the molecular weight of the intact molecule indicated that DNA was made up of two chains, as predicted by the Watson Crick model, and was not some multichain entity.
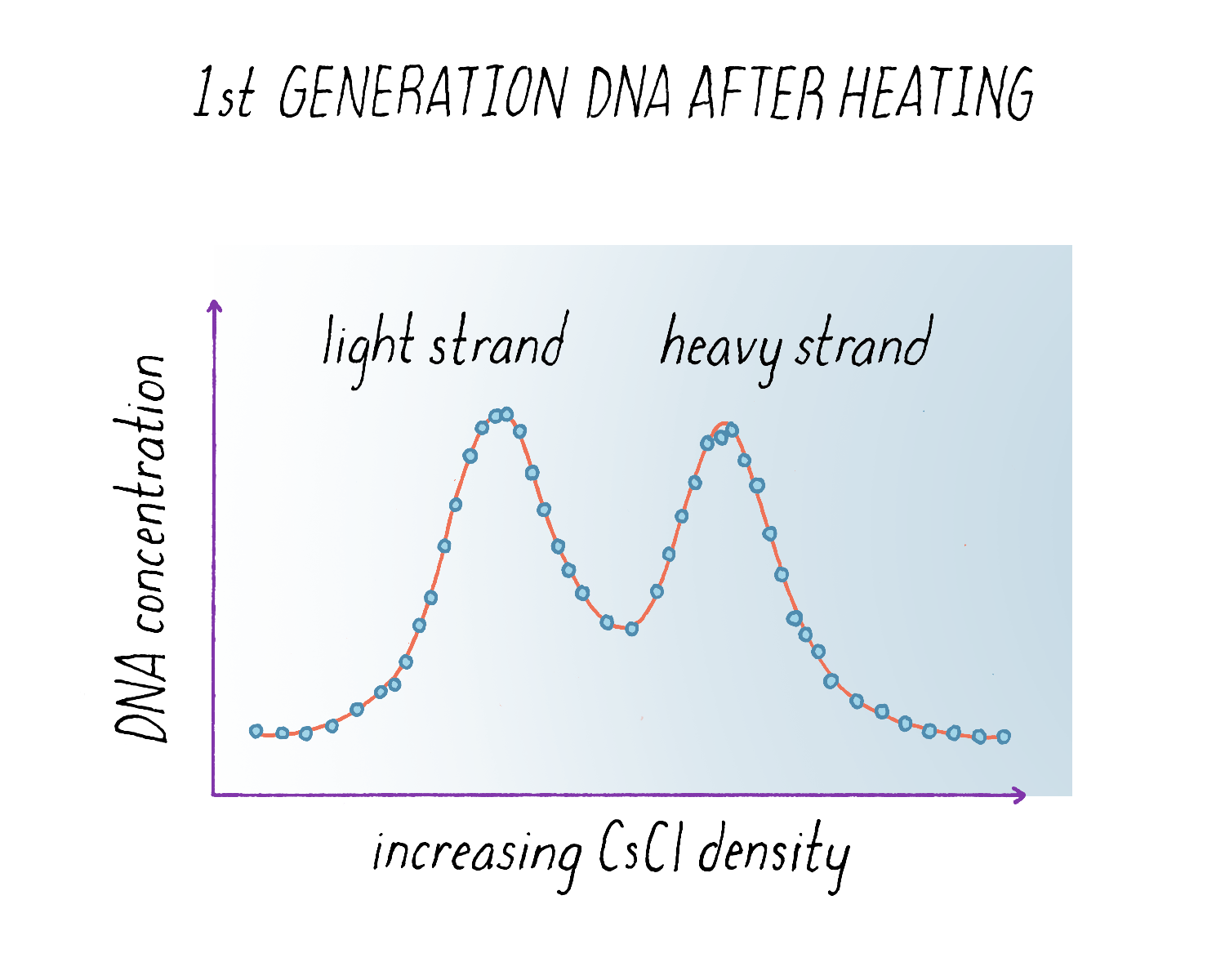
Based upon the results in Figure 9 and Figure 12, we concluded that:
1) The nitrogen of a DNA molecule is divided equally between two subunits. The subunits remain intact through many generations.
2) Following replication, each daughter molecule receives one parental subunit and one newly synthesized subunit.
3) The replicative act results in a molecular doubling.
These conclusions precisely aligned with the Watson–Crick Semi-Conservative model for DNA replication. DNA, as a double-stranded helix, unwinds, and each strand serves as a template for the synthesis of a new strand.
What Happened Next?
Frank and Matt
When we had our result, Matt quickly shared the news with Jim Watson in a letter dated November 8, 1957 (available for the first time here). It was common in those days to share results with colleagues through letters prior to a publication.
We also shared our results with Max Delbrück who took the news well that his Dispersive Replication model was incorrect. In fact, he wrote to a colleague that Meselson and Stahl had obtained a "world shaking result." But we were slow to get our work written up for publication. Once we knew the answer, we were keen to move onto new experiments rather than writing up our results.
Finally, Max had enough of our dallying and brought us down to the Caltech marine station at Corona del Mar. There, he quarantined us to a room in a tower, saying that we could not come out until we had written a draft of our paper. He was not being unkind, and we thought it great fun. Max's wife Manny Delbrück kindly came in occasionally to bring us delicious sandwiches, and Max also kept us company. We worked for 2 days straight and got him a draft.
Shortly thereafter, we completed our paper and Max communicated it in May 1958 to the Proceedings of the National Academies of Science, 4 years after our meeting at the Marine Biological Laboratory but less than a year after finally getting our experiments to work.
After our paper was published, we went separate ways in our lives. Frank got a job at the University of Missouri but soon thereafter moved to the University of Oregon in Eugene. Matt got promoted from a postdoctoral fellow to an assistant professor at Caltech and was teaching physical chemistry. However, the constant teaching limited time in the laboratory. Matt asked to be demoted from assistant professor back to senior postdoc, so he could get more work done in the lab. This is perhaps the only case in the history of Caltech in which a professor asked to move down the academic ladder. After a year as a senior postdoc, Matt then moved to Harvard to become an associate professor.
Decades have passed, and we now know much about the machinery that orchestrates DNA replication, including the unwinding of the strands and the synthesis of a new strand from the parental template. The details are beyond what can be discussed here, but you can view an animation of this process in Video 2 .
Closing Thoughts
When we first discussed the use of density labeling to test models for DNA replication under the gin and tonic tree, we could not imagine the psychological effect our experiment would have on the field. Many scientists were not initially convinced by the Watson–Crick model for the structure of DNA or their proposal for its mode of replication. It was not clear if their model could explain heredity and the properties of genes. Some people seemed to think the model was too simple to be the gene. Others thought it too simple (meaning too beautiful) to be wrong!
However, after our experiment, the DNA model seemed very real. We could watch DNA with a camera; the visualization of DNA bands was simple and clear. Our results showed that the gene is made of two complementary halves, each a template for the other. Even the disbelievers, such as the deeply thoughtful Max Delbrück, acquiesced. DNA was no longer an imaginary molecule in the heads of Watson and Crick. It was a dynamic molecule; one could perform experiments on it, and it behaved in living cells as one might predict. Mendel's concept of a discrete "factor" that could determine a plant character and remain intact generation after generation and the physical reality of a gene as double-stranded DNA became intertwined from that moment on.
It is gratifying to think that our experiment, so simple by modern standards, is still valued and taught. But beyond the logic of how the experiment was performed, we hope that our story also conveys other important lessons about science.
• Every hypothesis needs to be rigorously tested with a clear experiment.
• An atmosphere of freedom is important. We were both very junior at the time of this experiment, but we were supported by senior scientists who encouraged us to pursue our own ideas.
• Success does not come immediately. Reading most scientific papers (including ours), everything seems straightforward and works right away. Our narrative shows that the so-called "most beautiful experiment in biology" had some unsuccessful excursions and two years of work to come to successful finish.
• Because success does not come immediately, it is valuable to be able to share difficult times with a friend. We kept each from getting discouraged. There was a certain gaiety in our work. We even had fun when things went wrong.
• Much of science is built upon collaboration and friendship. This Key Experiment could never have been the "Meselson Experiment" or the "Stahl Experiment." The "Meselson and Stahl Experiment" required both parties. We complemented each other scientifically and encouraged each other personally. Well more than a half-century has passed since this experiment was performed, and we remain good friends today.
Dig Deeper
Dig Deeper 1: Alternatives to Semi-Conservative replication
Max Delbrück, in his 1954 paper (PNAS 40: 783-788), said the following of the Watson and Crick Semi-Conservative replication mechanism:
"The principal difficulty of this mechanism lies in the fact that the two chains are wound around each other in a large number of turns and that, therefore, the daughter duplexes generated by the process just outlined are wound around each other with an equally large number of turns. There are three ways of separating the daughter duplexes: (a) by slipping them past each other longitudinally; (b) by unwinding the two duplexes from each other; (c) by breaks and reunions. We reject the first two possibilities as too inelegant to be efficient and propose to analyze the third possibility."
Max's solution was to break the single chains at regular intervals, allowing rotation about single bonds of the unbroken chain followed by joining in a way that dispersed short segments of parental DNA among the single chains of the daughter duplexes. This is the Dispersive Model presented in Figure 5 and presented in more detail in Figure DD1. There was a germ of truth in Delbrück's idea of breakage. We now know of topoisomerases, enzymes that facilitate DNA replication by temporarily breaking, allowing unwinding, and then rejoining DNA. There are also enzymes that unwind DNA helixes called DNA helicases. Both enzymes use chemical energy derived from hydrolyzing adenosine triphosphate to perform work on the stable DNA double helix.
Another type of solution to the "unwinding problem," one that required no breaking and no entangling of the daughter molecules, was to imagine that the synthesis process would cause the entire parental duplex to rotate one turn for each turn of DNA synthesized. But this posed problems of its own – giving rise to a variety of long-forgotten proposed models, including evoking a motor at the growing point that would drive the rotation of the parent molecule, as proposed by John Cairns and Cedric Davern [J. Cellular Physiology, 70: S65–76 (1967)].
Alternative solutions questioned the DNA double helix model, but not semi-conservative replication. For example, one idea was to assume that the two chains are not wound around a common axis, but instead are simply pushed together (plectonemic coiling), which would require no unwinding and no rotation. This possibility, although it appeared remote, was not rigorously ruled out until much later in a paper by Crick, Wang, and Bauer in 1979 (J. Mol. Biol, 129: 449–461).
Dig Deeper 2: The idea for using density for separation
Matt
The idea for using density as a separation method came to me early in 1954 while I was a first-year graduate student at Caltech listening to a lecture by the great French scientist Jacques Monod. Monod was describing the problem of regulation of an enzyme called beta-galactosidase. If the bacteria were growing in a medium without lactose (a sugar), the enzyme activity was very low. When lactose or a chemical analogue of lactose was added, the enzyme activity was induced. The question was how? One model was that the enzyme was always there, but is inactive unless lactose is around. Another model (the correct one) was that the enzyme is synthesized de novo after the inducer is added. I thought that it might be possible to measure new enzyme and distinguish it from old enzyme if the enzyme was synthesized from heavier building blocks (amino acids). How could one make heavier building blocks? I thought that deuterium (a heavy isotope of hydrogen; 2H) might be the answer. If one grew bacteria in heavy water (2H2O) and switched to normal water (1H2O) when one added inducer, then any newly synthesized beta-galactosidase would have had a greater density than the pre-existing beta-galactosidase. I never did the experiment, but the idea primed me for the DNA replication problem.
Dig Deeper 3: The role of the centrifuge in the Meselson–Stahl experiment
Matt describes briefly how this technique evolved
The first paper (see the reference list) that Frank and I wrote together (along with Jerome Vinograd) was on the method and theory of using centrifugation in an equilibrium density gradient, which showed that this method not only could separate molecules but could also be used as a tool to determine their molecular weights. This work was also part of my PhD thesis at Caltech. When I presented this work at my thesis defense, the great physicist Richard Feynman was on the examination committee, along with Pauling, Vinograd, and one of Pauling's post-docs who taught me X-ray crystallography. Feynman had not read the thesis but did so during the defense. I presented my rather long mathematical derivation showing that macromolecules in a density gradient in a centrifugal field would be distributed in a Gaussian manner about the position of neutral buoyancy with the width dependent upon the square root of the molecular weight. Feynman then went to the blackboard and, on the spot, produced a much shorter derivation of the same thing, modeled on the wave function for the quantum mechanical harmonic oscillator. Feynman writes about our experiment in his jolly book, Surely You're Joking, Mr. Feynman.
The way in which we found that CsCl forms a density gradient on its own was somewhat fortuitous. We initially thought that we needed to pour a CsCl gradient in the tube in advance. However, we found that just by centrifuging an initially homogeneous solution of cesium chloride produced a continuous gradient density on its own after several hours. From the width of the DNA band in the density gradient, we could also calculate the molecular weight of the DNA molecules in the gradient as 7 million Daltons. The chromosome of E. coli is much, much larger, but long molecules of DNA are fragile and had been broken up by shear forces while passing through the hypodermic needle with which we loaded the centrifuge cell. Subsequent to our result, CsCl equilibrium density-gradient centrifugation became a standard tool for isolating DNA from cells for decades and was used in important experiments such as the demonstration of messenger RNA by Brenner, Meselson, and Jacob and showing the mechanism of general recombination in phage lambda by Frank Stahl.
References and Resources
Guided Paper
Meselson, M. and Stahl, F.W. (1958). The replication of DNA in Escherichia coli. Proceedings of the National Academy of Sciences U.S.A., 44: 672–682.
This classic paper provides experimental evidence that the strands of the DNA double helix serve as templates to create a new copy of DNA. These results provide experimental evidence of The Watson and Crick model of DNA replication (‘semi-conservative replication’) demonstrating that genetic information is passed from one cell or organism to its progeny.
DownloadReferences
- Matthew Meselson’s letter to James Watson from November 8, 1957, describing the results of their experiments on DNA replication. Download.
- Meselson, M., Stahl, F.W., and Vinograd, J. (1957). Equilibrium sedimentation of macromolecules in density gradients. Proceedings of the National Academy of Sciences U.S.A., 43: 581–588.
This paper describes the use of the centrifuge and density gradient to analyze biological molecules, a technique that was used in their 1958 paper but also very broadly used for many applications in biology. See also Dig Deeper 3.
- Holmes, F. L. “Meselson, Stahl, and the Replication of DNA. A History of ‘The Most Beautiful Experiment in Biology’” (2001). Yale University Press
An outstanding resource for those wanting a detailed, accurate description of the Meselson–Stahl experiment.
Resources
- White Board Video on the Semi-Conservative Model of DNA and the Meselson–Stahl Experiment by iBiology. https://www.ibiology.org/genetics-and-gene-regulation/semi-conservative-replication-of-dna/
A nice 7:30 min video describing the Meselson–Stahl experiment and its conclusions.
- The Double Helix: https://www.hhmi.org/biointeractive/double-helix
This film documents the discovery of the structure and replication of DNA including interviews with James Watson who, along with Crick, proposed the double helix model of DNA.
- Pulse Chase Primer: The Meselson–Stahl Experiment: https://www.hhmi.org/biointeractive/pulse-chase-primer-meselson-stahl-experiment
This activity is often used in conjunction with the short film The Double Helix. It introduces students to Meselson and Stahl experiment and helps them understand the concepts generated via those experimental results.
- HHMI Biointeractive DNA Collection: https://www.hhmi.org/biointeractive/dna-collection
This collection of resources from HHMI Biointeractive addresses many of the major concepts surrounding DNA and its production, reading, and replication.